The data you need already exists. Getting to it is another story.
Every collaboration starts with a data access agreement. Every agreement triggers legal, ethics, IT security.
Months pass — or it never happens. tracebloc removes data transfer and direct access — the rest is routine.
How It Works
Your model travels to the data. Not the other way around.
You submit
You get results
The problem
The data exists. The ecosystem refuses to expose it.
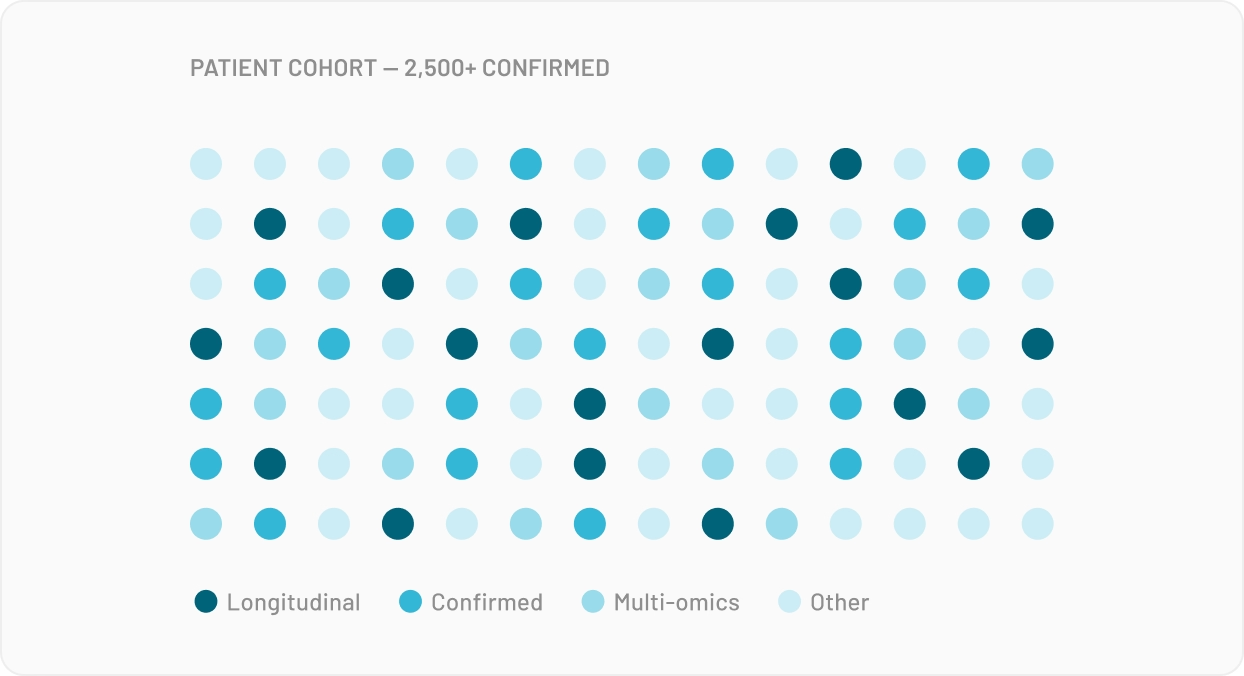
The cohort your reviewer wants exists somewhere, but you can't get to it in time. An MTA takes eight months, an Epic integration costs $25k before any line of code is written. The revision deadline is long gone before access can be set up.
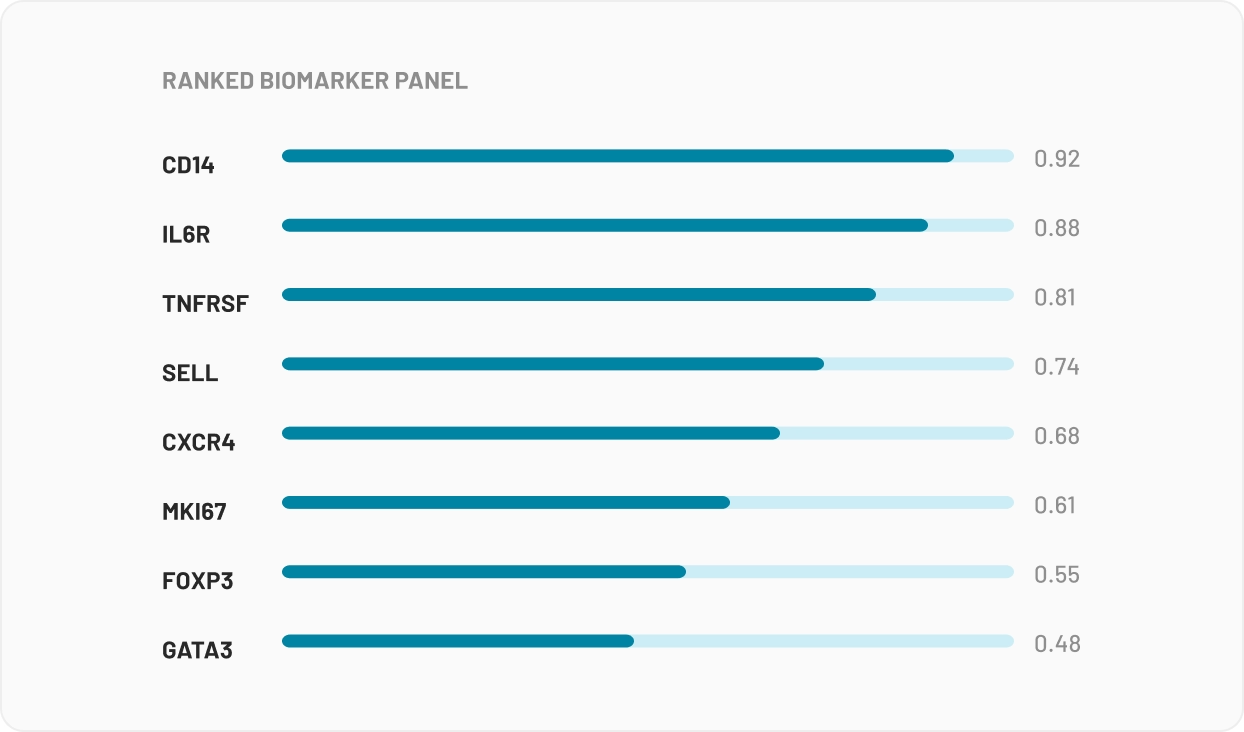
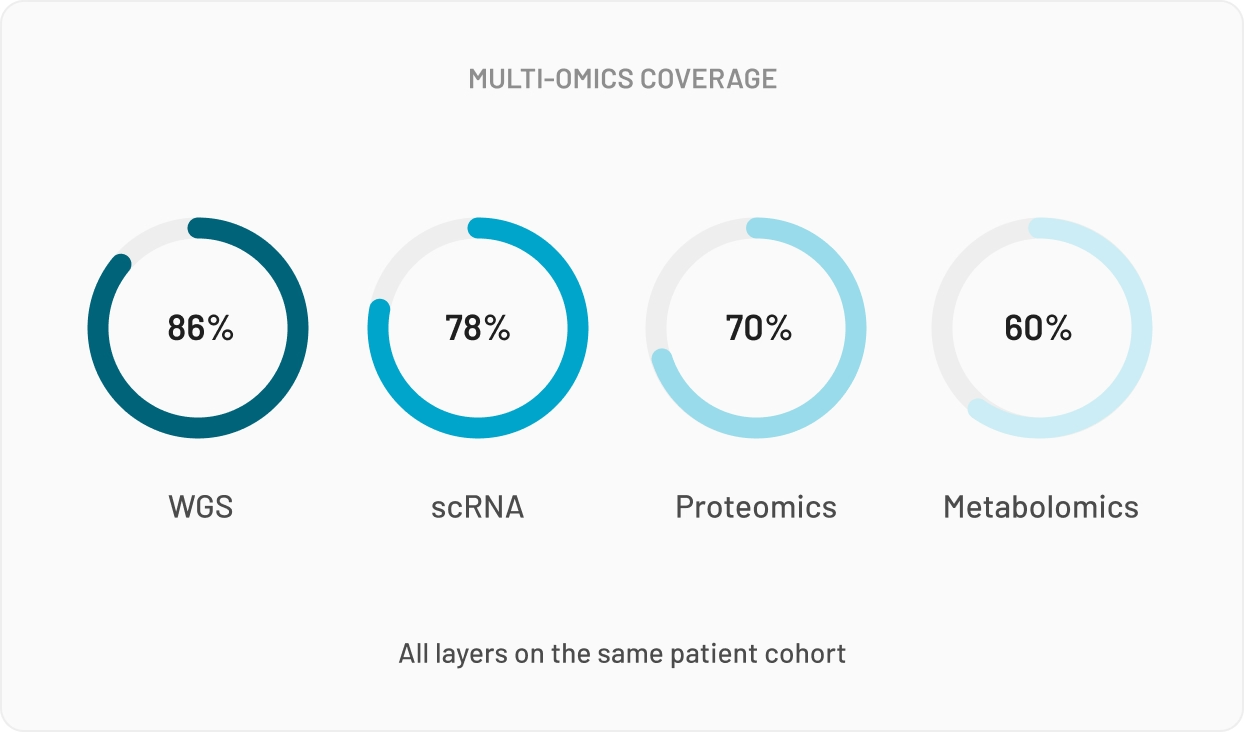
For Multi-Omics data, your n is typically small, and so is everyone else's in your network. Multi-omics needs numbers no single site can produce, which is why bioinformaticians keep asking whether they can combine datasets from different studies. The answer: statistically yes, legally no.
tracebloc is what you give your data partner so they can say yes. They deploy a workspace on their infrastructure. You submit your model. No data leaves their site — so the approval that used to take months is now routine.
The Platform